Biology
The Biology Department has a DNA Sequencing Core Facility, servicing several CSAM Departments
This state-of-the-art instrument is used worldwide for genetic sequencing and fragment analysis (AFLP, SNP, and microsatellite analysis)
Common applications include:
- DNA fingerprinting in crime labs
- identification of genetic diseases in hospitals
- identification of horses for thoroughbred breeding and racing
This facility has been used to support both work of individual laboratories as well as sequencing projects in undergraduate teaching laboratories
Examples of recent projects:
- human mitochondrial D-loop analysis
- identification of fungal pathogens in amphibians
- bacterial metagenomic analysis in soil and water samples using the 16S locus
- genetic barcoding of jellyfish from Barnegat Bay

This inverted microscope is set-up to measure bioelectrical currents and study the activity of specialized membrane proteins, called ion channels, using a technique known as patch clamping.
Usage:
- Glass pipettes are fabricated with tips only a few micrometers in diameter to be directed towards the cell using a micromanipulator until it makes contact with the cell membrane.
- The cell membrane forms a tight seal with the pipette glass and can be ruptured by applying suction opening the electrical environment inside the cell to an electrode housed higher in the pipette.
- An ionic solution fills the pipette allowing the control of the ionic environment both inside and outside the cell.
- To maintain mechanical stability, the microscope is housed on an air-suspended vibration isolation table.
Since the currents are often on the order of one billionth of an ampere, the set-up is protected from outside electrical noise by a housing of conductive material, known as a Faraday cage.
This technique can be combined with fluorescence microscopy techniques, such as calcium imaging, in order to tease out the activity of different ion channels.

Used for tissue culture work
Enables one to see living growing cells
Currently, the equipment is being used to study the effects of the dust from the World Trade Center 9-11 attack on human lung cells

- For quantification of DNA (or RNA)
- PCR (Polymerase Chain Reaction)
- Provides a very accurate determination of DNA copy number in samples
- Great when looking at relative abundance/diversity of species or gene expression
- Used frequently for both research and teaching laboratories
- Capable of performing both SYBR-Green and Taqman-based assays
- Has four channels permitting multiplexing of samples
- Examples of recent projects:
- microbial source tracking of nonpoint source pollution in New Jersey rivers
- temporal and spatial distribution of sea nettle (Chrysaora quinquecirrha) DNA in Barnegat Bay
- detection of fungal pathogens in amphibian populations
- Department taught a laboratory methods class (BIOL598) on quantitative PCR where students were able to quantify microbial contamination of food and environmental samples

Computer Science
Used in the Computational Sensing Laboratory managed by Dr. Stefan Robila.
Hyperspectral imagery (HSI) collects data as hundreds of images (spectral images or spectral bands), with each image corresponding to narrow contiguous wavelength intervals.
Hyperspectral sensors cover wavelengths from the visible range (0.4μm-0.7μm) to the middle infrared range (2.4μm).
The ability to distinguish minute differences between materials’ spectral properties lead to HSI based solutions in agriculture, mining, environmental sciences, food production, etc.
Introduced only a few decades ago, it is clear that HSI is now a well-established tool that has been used here for a variety of projects including face detection and recognition, real-time collection and processing of data, as well as spectral unmixing.
Students at all levels (graduate, undergraduate and high school) with interests in computing, biology and environmental sciences have the opportunity to design and implement experiments using the camera.

Refers to computational environments comprised of tens to hundreds of thousands of computing units, all working together and delivering extraordinarily large computational power.
Montclair State hosts several HPC environments including two as part of the Computational Sensing Laboratory led by Dr. Stefan Robila, professor of Computer Science.
The first HPC system is a computer cluster formed of a master node and (64) compute nodes. Each node consists of two, quad-core 2.4GHz processors, 16GB of RAM and 50GB of storage. In total, the cluster provides (520) 64-bit, 2.4GHz cores, 1.008TB of RAM and 3.7TB of storage. Programming on the cluster can be done in multiple languages and development environments including MPI and OpenMP.
The second HPC system is formed of a Microway computer equipped with a Graphics Processing Unit (GPU). The GPU, part of the NVIDIA Tesla series has 2,496 computing cores and a total of 5GB on chip memory.
Both systems were extensively used in projects by faculty and students in the departments of Computer Science, Biology, Linguistics and Mathematics.

Earth and Environmental Studies
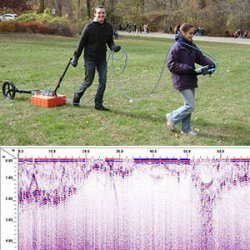
Allows for the investigation of the shallow subsurface.
Antenna emits electromagnetic waves into the ground and records wave reflections.
Lower frequency antennas (e.g., 100 MHz) can penetrate tens of meters into the ground.
Higher frequency antennas (e.g., 900 MHz) can only penetrate tens of centimeters into the ground and detect much smaller objects and finer features.
We have: 100 MHz and 270 MHz, which are most useful for geologic and environmental applications, such as mapping sediment layers, imaging shallow faults and sometimes detecting groundwater.
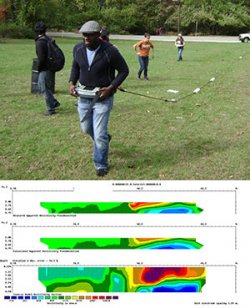
Used for resistivity profiling.
Electrical current is put into the ground via capacitors and the voltage drop between the transmitter and receiver is measured to determine the resistivity of the ground using Ohm’s Law: voltage = current * resistance.
Increasing the spacing between the transmitter and receiver increases the depth of penetration of the electrical current and thus increases the depth of measurement.
Our configuration includes two receivers and one transmitter so that we can collect data at two depths with each pass.
Resistivity profiling is commonly used to map groundwater because the presence of water in soil or rock pores greatly reduces its resistivity.
Using a sledgehammer and an aluminum plate as our source, we can send seismic waves into the subsurface to investigate shallow features like faults, sediment layers and depth to bedrock.
Our seismic system consists of 48 geophones to detect the seismic waves, two 24-channel Geometrics Geode seismometers to record and digitize the geode signals, and a rugged field laptop to control the system and process the data.
In seismic refraction surveying, the geophones are laid out in a line and the sledgehammer is used at several shot points along that line. In most places, the waves will be refracted – or bent – where they reach an interface between materials with different wave speed, for example, between soil and bedrock, similar to how light is refracted at the interface between air and water.
Arrival times of the first waves to reach the geophones are used to determine the depth of such interfaces below our survey line

Surveying instrument that allows for site surveys and in-field measurements of river channels, hillslopes, and other topographic features
Has been used by our students to measure lake bathymetry, river channel dimensions and erosion on hillslopes
The precision is ±1mm
Great tool for measuring the Earth’s surface to high levels of precision

Measures small variations in the magnetic field as it is carried over the ground
Materials with high magnetic susceptibility (e.g., ore bodies, buried metal objects) will produce a positive magnetic anomaly that the magnetometer can detect
Parallel survey lines over an area allow us to create maps of the magnetic anomalies

